The path from concept to a clinically effective medicine is long and expensive. Many drug candidates fail not only because the chosen target is incorrect, but also because their pharmacological effects, safety profiles, or pharmacokinetic properties are not fully understood until late in development. In this context, in vitro pharmacology has become a central pillar of modern research, providing controlled, high-content data on how molecules interact with biological systems long before they are tested in humans.
What Is In Vitro Pharmacology?
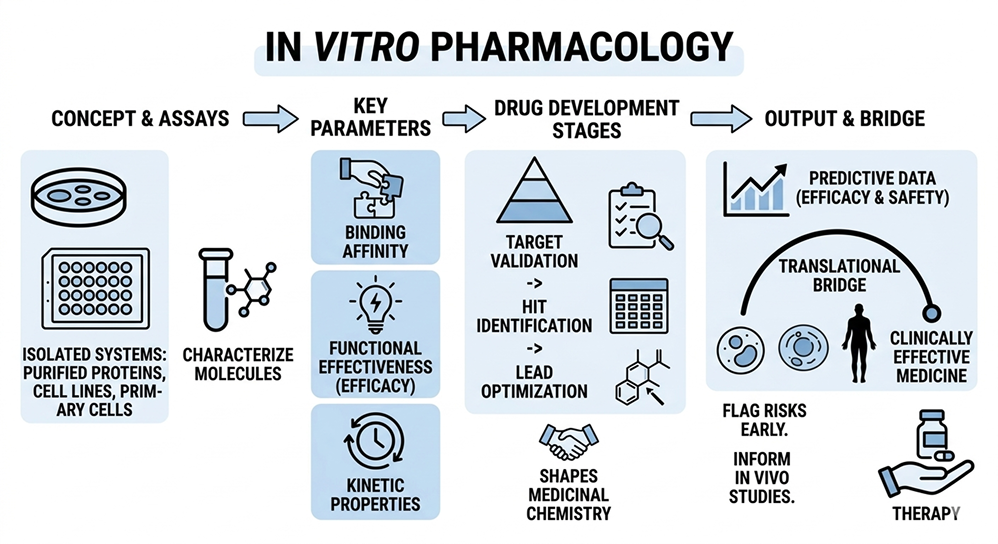
In vitro pharmacology studies drug action in isolated biological systems outside the living organism. Using purified proteins, recombinant receptors, cell lines, primary cells, or tissue preparations under controlled conditions, researchers characterize compound–target interactions. Key parameters include binding affinity, functional efficacy, kinetic properties, and modulation of signaling pathways – all assessed without whole-body physiological interference.
The term "in vitro" contrasts with in vivo pharmacology, which evaluates effects in intact animals or humans and accounts for absorption, metabolism, and organ cross-talk. This reductionist approach – rooted in early biochemistry of purified enzymes – has progressed from basic protein assays to complex 3D organoids, unified by experimental controls that focus on specific molecular or cellular components.
Spanning target-based assays (direct molecular interactions) and phenotypic assays (cellular behavior changes), in vitro assays bridge basic disease biology and clinical reality, enabling the study of disease pharmacology in vitro and in vivo, supporting medicinal chemistry efforts, and shaping follow-up in vivo experiments.
The Role of In Vitro Pharmacology in Drug Development
The role of in vitro pharmacology in drug development extends across all key stages:
Target Validation. Before investing in compound screening, researchers must confirm that a biological target is relevant to disease pathology. In vitro studies allow researchers to assess how modulation of a specific receptor, enzyme, or pathway elicits a measurable, disease-relevant cellular response. Through genetic manipulation techniques such as CRISPR or RNA interference, researchers can validate whether altering target expression affects disease models at the cellular level in vitro.
Hit Identification. High-throughput in vitro pharmacology screening is used to test large compound libraries. These assays measure binding affinity, functional activation, or inhibition, allowing rapid identification of initial hits. Automation and miniaturization have dramatically increased screening efficiency while reducing reagent costs and variability.
Hit-to-Lead and Lead Optimization. Once hits are identified, in vitro pharmacology becomes even more critical. Medicinal chemists rely on detailed potency, selectivity, and mechanistic data to refine compounds. Assays evaluating receptor subtype selectivity, signaling bias, metabolic stability, and cytotoxicity guide chemical modifications. This stage represents a constant dialogue between chemistry and biology, where in vitro pharmacology techniques provide the experimental evidence that shapes compound optimization.
Translational Bridge to In Vivo Studies. In vivo pharmacology ultimately assesses efficacy and safety in whole organisms; robust in vitro data significantly reduce risk. Well-designed in vitro pharmacology assays help prioritize candidates with favorable profiles before moving into animal models.
Core In Vitro Pharmacology Techniques
The in vitro pharmacology toolbox provides a diverse set of experimental approaches that enable researchers to investigate drug action at multiple levels of biological complexity.
At the molecular level, biochemical assays evaluate direct compound–protein interactions, including enzyme inhibition, radioligand binding, and fluorescence-based readouts. These methods enable rapid screening of large libraries and generate clear structure–activity relationships, although they do not fully capture cellular context.
At the cellular level, compounds are assessed in living systems. Reporter gene formats track transcriptional responses, while phenotypic assays monitor behaviors such as proliferation, migration, differentiation, or cell death. High-content imaging further supports automated analysis of multiple cellular parameters simultaneously.
Advanced three-dimensional culture systems better preserve tissue organization and cell differentiation, often demonstrating greater predictive power than traditional 2D cultures. Organ-on-chip platforms incorporate microfluidic systems that mimic blood flow and mechanical forces, enabling studies under conditions that approximate human physiology.
Throughout these techniques, comprehensive assay validation supported by robust quality control frameworks is essential to generate data that guide compound progression decisions.
Integrating Machine Learning & Data Science in In Vitro Studies
As in vitro pharmacology has become more sophisticated, the volume and complexity of data generated have grown dramatically. High-content imaging, multiplexed biomarkers, and transcriptomic readouts can produce thousands of parameters per condition, generating datasets that challenge traditional statistical analysis. Machine learning and data science tools are increasingly used to extract patterns from these rich datasets, linking chemical structure to in vitro pharmacology assays and, ultimately, to in vivo pharmacology outcomes. This enables better prediction of efficacy, safety, and translational potential.
In practical terms, data-driven models can guide the design of new analogs, prioritize which molecules to test experimentally, and flag unexpected off-target effects. Used across coordinated research efforts, these approaches promote the sharing of insights across different research projects and the continuous improvement of predictive models. Importantly, such models do not replace experiments but help focus in vitro pharmacology on the most informative assays and conditions, making better use of limited resources.
Benefits and Challenges of In Vitro Pharmacology in Drug Development
In vitro pharmacology offers distinct advantages for modern drug discovery. By design, these assays can be run rapidly, adapted to high-throughput formats, and maintained under stringent experimental control, enabling the systematic exploration of diverse chemical entities and biological responses while keeping resource use manageable. They enable systematic investigation of structure–activity relationships, flag potential safety liabilities at an early stage, and dissect mechanisms of action with a level of detail that would be difficult or ethically problematic in animals or humans. When thoughtfully implemented, in vitro pharmacology techniques also contribute to the 3Rs framework by limiting, and in some cases replacing, the need for animal use.
However, in vitro experimental models have limitations. Simplified systems may fail to capture key aspects of disease biology relevant to in vivo contexts, including immune interactions, complex tissue architecture, and whole-body pharmacokinetics. Results can be heavily model-dependent: a compound that looks promising in a single cell line may underperform in primary cells or fail in vivo. Translational limitations may stem from biological divergence between human and animal systems or from oversimplified culture conditions. Consequently, even in vitro assays require contextual interpretation alongside complementary datasets from in vivo pharmacology and clinical literature.
Future Trends and Research Directions in In Vitro Pharmacology
In vitro pharmacology is advancing through rapid technological innovation and increasing emphasis on human-relevant experimental systems. Microphysiological systems and organ-on-chip platforms are transitioning from experimental concepts into practical screening tools, enabling the recreation of key features of human organ function in miniaturized, testable formats.
At the same time, patient-derived models such as organoids and induced pluripotent stem cell systems allow researchers to study drug responses within specific genetic backgrounds, supporting the move toward precision medicine. The integration of multi-omics approaches further strengthens this capability by revealing how compounds influence biological systems at the transcriptomic, proteomic, and metabolic levels, while machine learning helps interpret these complex datasets.
Automation and miniaturization are expanding throughput, particularly in phenotypic screening methods that capture compound-driven biological effects at the cellular level without predefined targets. Increasingly, these experimental approaches are combined with computational tools – including molecular dynamics, structure-based design, and AI-driven prediction – forming integrated in silico and in vitro workflows.
Regulatory agencies are also beginning to recognize the value of in vitro data as partial alternatives to animal testing, especially in safety assessment. Together, these developments are shaping future platforms that deliver reproducible, physiologically relevant insights while remaining compatible with the speed and scale required in early drug discovery.
Thus, as drug development grows increasingly complex, the thoughtful application of in vitro studies in drug discovery – supported by interdisciplinary expertise and integrated drug discovery services – will remain essential to turn scientific insights into effective therapies.